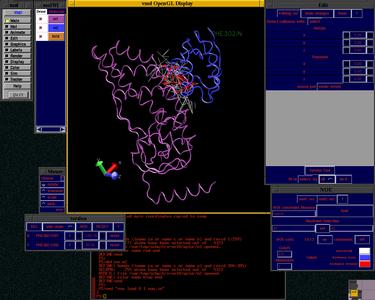
vmd-xplor NMR Visualization
The vmd-xplor package is a combination of the X-PLOR structure determination program and VMD (Visual Molecular Dynamics) - a freely available molecular visualization program. This package allows manual manipulation of the protein structures to satisfy experimental NMR data, and can also be used to visualize the goodness of fit of a particular model to given restraints.
Reference:
| Download . |
VMD-XPLOR Tutorial - Example walkthroughs. |
current version: 1.17
Version 1.14 or higher is required when using with Xplor-NIH version 3.0 or newer.
Note: For versions 1.2 and newer, you must download xplor-nih separately, and specify its location during installation of vmd-xplor.
- based on VMD-1.9.3.
- molWid now supports multiple representations and grouping structures for group operations (e.g. setting the color of a subset of structures)
- better support for multi-monitor setups
- improved keyboard shortcuts.
- noeWid now supports ASSIgn ... OR syntax and center averaging
- by default, each model in PDB files are now loaded as separate structures
- other fixes and improvements.
- Intel/Linux - support for kernels 2.4 and newer for x86 and amd64 platforms.
- MacOS X (under an X server).
- language extensions: small module added.
- remote execution of X-PLOR. Threaded for simultaneous calculation and communication.
- communicates molecular structures, animation sequences, etc. to VMD via the tcl-DP sockets package.
- X-PLOR is written in Fortran, extensions in C++.
- easy-to-use molecule display list
- toggling molecule on/off
- quickly focus on molecule of interest.
- change molecule properties
- allows large number of simultaneous structures
- edit tool
- rotate, translate one structure relative to another
- fit to another structure
- change torsion angles
- detect collisions with other molecule
- display NOE restraint information as lines.
- dynamically updated as molecule is edited.
- allows inter- or intra-molecular NOEs
- select a subset of restraints to view
- text annotations displayable
- text view of violations
- display torsion-angle restraints - same functionality as for NOE info
- used TCL scripting and C++ back-end where necessary. TCL-Tk: fast GUI development.
Thanks to: summer student Devtulya Kavathekar for his work on the most recent release of VMD-XPLOR.
question/comments? Contact charles.schwieters@nih.gov .